Reaching the exabyte level
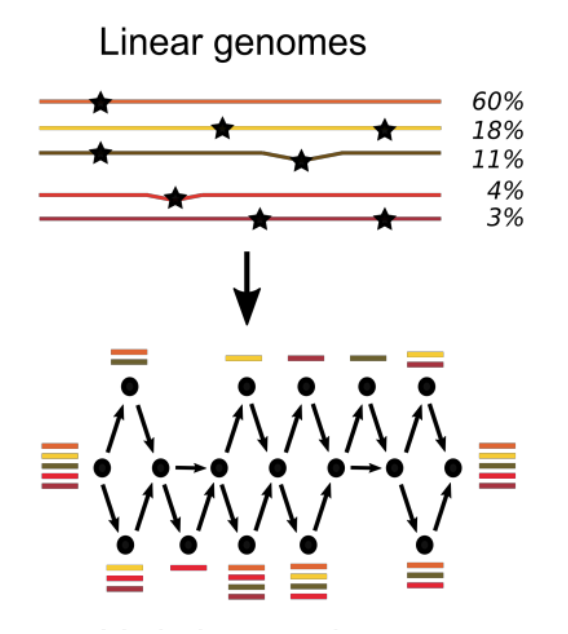
The aim of the ALPACA project will be to develop new algorithmic approaches with the capacity to represent genomes in graph form, as opposed to "flat" sequences.
The project deals with the vast quantities of data generated by sequencers. “We’ve reached the exabyte level. This means we have to find another way of storing and processing information.” explains Pierre Peterlongo, the man in charge of the GenScale project team at the University of Rennes Inria centre. This data takes the form of series of letters (A, C, G, T) corresponding to nucleotides, the building blocks of DNA. These letters are currently stored sequentially, one after the other, in long text files.
"It is currently quite common to sequence multiple individuals belonging to the same species. There was the 1,000 Genomes project, which catalogued genetic variations in humans. A 100,000 Genomes project has just been launched. But to use you and I as an example, our genomes are 99% identical. It therefore seems a shame to store information as redundant as this practically twice in two files of text sequences which are so similar to each other. Designed to avoid such an occurrence, the concept of pangenomics involves organising information in graph form. Shared information is stored only once, while variants are represented by branches in a graph. This really helps to cut down on both space and redundancy."
And that’s not all. “These types of structures can also be used to derive biological information. Out of 1,000 individuals, let’s imagine that some of them have cancer. We will be able to look at a graph and identify certain areas more specific to this disease, thus allowing us to develop a better understanding of the mechanisms involved. Such variants might seem to play a role, for instance, even if we don’t yet know how, and these are the ones we’ll look at. Graphs are a useful tool for this sort of work.”
An enriched data structure
However, human genomes are comprised of some three billion characters, meaning that visual navigation within a graph is not the best-suited method. “Statistical analysis is needed in order to identify areas of interest.” Scientists are also keen to re-insert any annotations which may have been made during research on the genome into these graphs. What’s more, “an enriched data structure makes it possible to record useful information.”
Inria and the Pasteur Institute have been tasked specifically with creating this graph-based structure. “We will likely develop a common tool, which we will then share with the whole community. We are also focusing on utilising this structure algorithmically.”
There are two main challenges. “The first relates to the mass of data. We must find a more effective way of storing information that will allow us to be more aggressive when it comes to compression, in addition to considerably speeding up graph queries. If you compress too much, it will take a long time to decompress before you are able to launch a query.” The issue lies in identifying the best compromise.
The second problem concerns updates. “Once you have built a graph of 1,000 individuals, when you add a 1,001st, you don’t want to have to start all over again from scratch. But we don’t yet have a data structure that is malleable enough to allow such easy updates to be made.”
A few steps further down the line from the ALPACA project, “other researchers are studying this new graph-based structure while working on their own data. My colleague Marie-France Sagot, a bioinformatician and head of the Erable project team at the Inria Lyon centre, works far more closely with biological data than I do. Her aim is to find solutions with the capacity to reveal more about specific types of data.”
The technology developed through this research is also targeted at industrial sequencers. “Their interest in it is twofold. Firstly, it will help to take the strain off their often saturated storage infrastructure. Secondly, they will be able to provide their clients with genomic data structured in a way that is more immediately usable.”
The type of EU funding
This is an "Innovative Training Network (ITN)", a Marie Sklodowska-Curie grant.
The project requires a range of skillsets in order for it to be a success, from pure algorithmics to data analysis and validation, and involves more than ten different EU partners. This type of funding is tailored to suit large-scale projects. ITNs are also designed to provide training to the next generation of researchers on the most important subjects of the future. Given the practical importance of biological data, it is vital for the next generation to engage with research into pangenomics.
“The idea behind this funding initiative is to bring together a range of European research institutes to tackle an emerging scientific problem, funding a large pool of PhD theses in order to bring through the next generation of researchers”, says Pierre Peterlongo, head of the GenScale project team at Inria centre at Rennes University. “Each student is assigned a supervisor within a laboratory, and a number of co-supervisors in other locations, including with an industrial partner, and will conduct part of their research at a partner laboratory. On top of this there are summer schools, seminars, and so on, ensuring young researchers have opportunities to properly get to know each other. This will eventually help to build a network, and will also free students from the somewhat restrictive framework of their subject of study, while familiarising them with other people's work, and enabling them to develop a more comprehensive understanding of the research topic.”
14 students of Computational Pangenomics will receive training through the ALPACA initiative between 2020 and 2023. Coordinated by Bielefeld University (Germany), the consortium is comprised of 23 academic and industrial partners, including: CWI (the Netherlands), the University of Pisa, the University of Milano-Bicocca (Italy), Heinrich Heine University Düsseldorf (Germany), Comenius University in Bratislava and Geneton (Slovakia), the University of Helsinki (Finland), the University of Cambridge (United Kingdom), Inria, the CNRS, and the Pasteur Institute (France) and the European Molecular Biology Laboratory (Germany). The total amount of funding is 3.67 million euros.