Could you give us a brief description of the project?
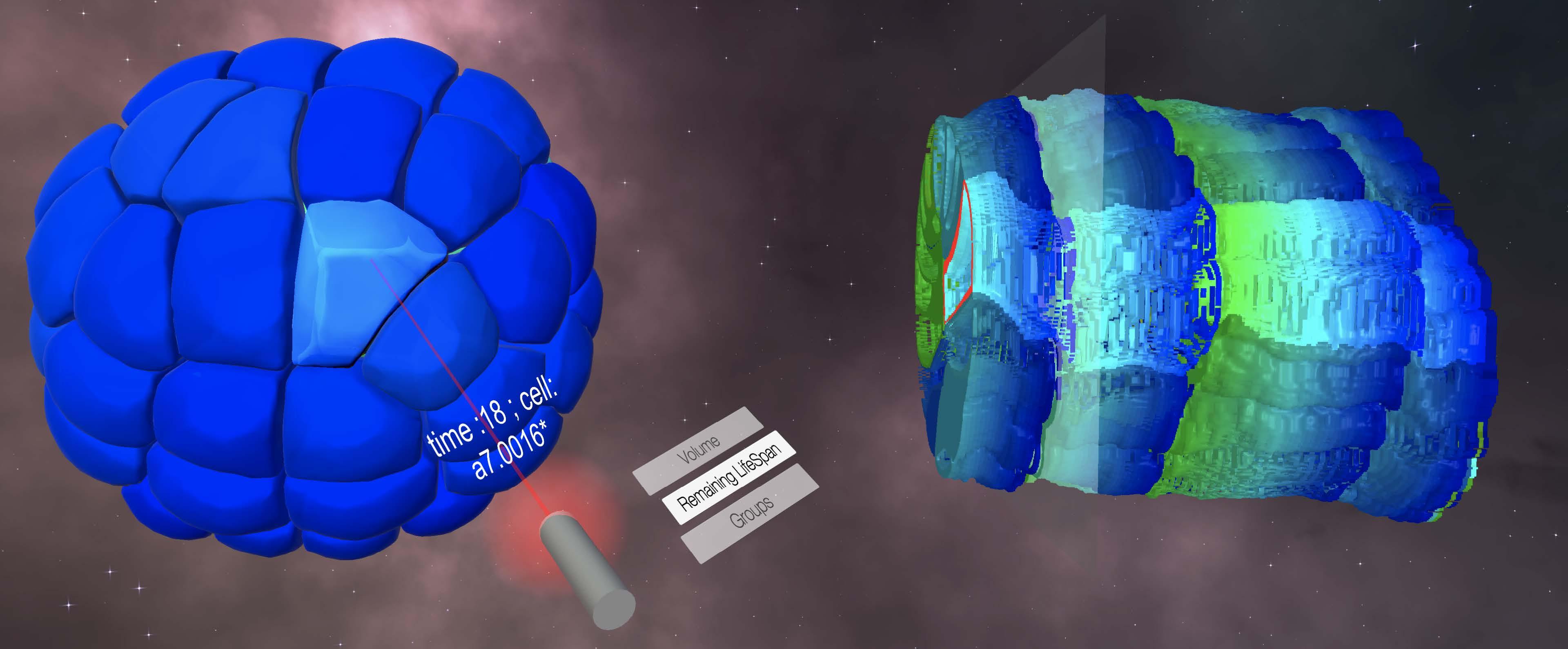
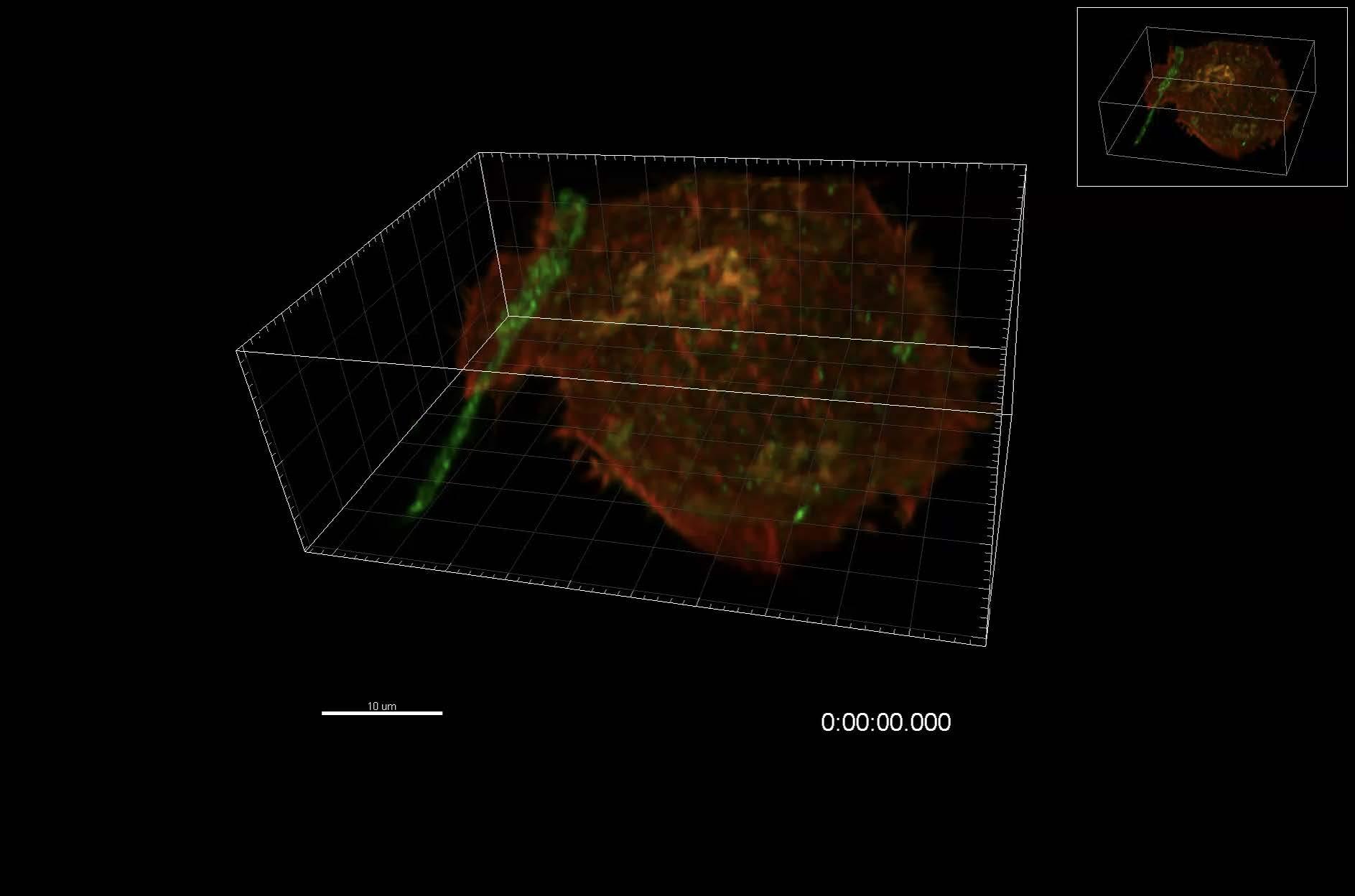
“The aim of Naviscope is to develop methodologies for use with the complex images encountered in microscopy, at both an intracellular level (inside cells) and a multicellular level (concerning groups of cells)”, explains Charles Kervrann, Naviscope Challenge coordinator. “At a spatial level, the sheer volume of data to be analysed can be overwhelming, making interpretation difficult.”
These are temporal series of 3D volumes, also known as 4D imaging (time being the fourth dimension)
The project’s objective is to develop software capable of interacting with and visualising complex 3D + time data, guided by learning via annotations stored in databases. This is something no software program is currently capable of doing on its own.
How did your project come about?
« Whenever we spoke with other partners acquiring images from 4D microscopy, a major issue they all seemed to be experiencing was working out how to navigate their way through all the data. »
The concept behind the Inria Challenges initiative is to try to put together joint projects, as opposed to projects devised by teams operating on their own. The main advantage of this is that it enables different project teams to pool and share their expertise: in the case of Naviscope, this was in « computer graphics, visualisation, machine learning, optics and cell biology. ».
« We develop 3D image processing techniques, but visualisation is not our team's (SERPICO) core skill. That’s how the collaborations with HYBRID and AVIZ came about, the goal being to benefit from their specialist skillsets. There is also an interest in it for these two teams who will be developing original methodological approaches linked to new application-based subjects and therefore leading to new opportunities »
What makes your project innovative?
Another aim of NAVISCOPE is to incorporate the principles of learning, which has been largely absent from conventional approaches to data visualisation. One way in which the project is original is through its aim of finding a way of designing a tool capable of guiding data exploration using learning techniques and to enable users to annotate the data.
What have the initial results been?
« The NAVISCOPE project was launched two years ago, and the results that we have been able to present are the fruit of one year of intense planning followed by a second year more geared towards practical application.»
We have already developed a way of enabling biologists to immerse themselves using a headset and to handle data by integrating derived data (digital or symbolic data), which was our first big breakthrough. NAVISCOPE has also been starting to produce results on questions surrounding data annotation, helping to build learning bases.
« In two years, the bulk of our objectives will be operational, and I believe we will have reached the proof of concept stage, i.e. when evaluations are carried out by beta testers (biologists) who will be able to certify whether the systems work or not. »
What will your discoveries be used for? Do you have any particular sectors in mind?
Our overarching objective is to enable biologists to make discoveries in biology using a tool such as ours.
« Another potential outcome might be the development of much simpler tools for teaching students of biology, letting them see molecules interacting with each other or to understand the dynamics of viruses within cells using a virtual reality headset, for example. »
What will the next steps be?
« We will be gathering together all of these software components into one single interoperability framework in order to make data manipulation simple and interactive, with online documentation. »
Coordinator : C. Kervrann.
Partners : The AVIZ project team (Saclay) ; The BEAGLE project team (Lyon), The HYBRID project team (Rennes), The MORPHEME project team (Sophia-Antipolis) ; The MOSAIC project team (Lyon), The PARIETAL project team (Saclay), The SERPICO project team (Rennes) ; The Unité MaIAGE INRAE (Jouy-en-Josas) ; The Institut Curie Paris (The STED and LOCCO project team) ; The Institut Pasteur Paris (The DBC project team) (Paris).